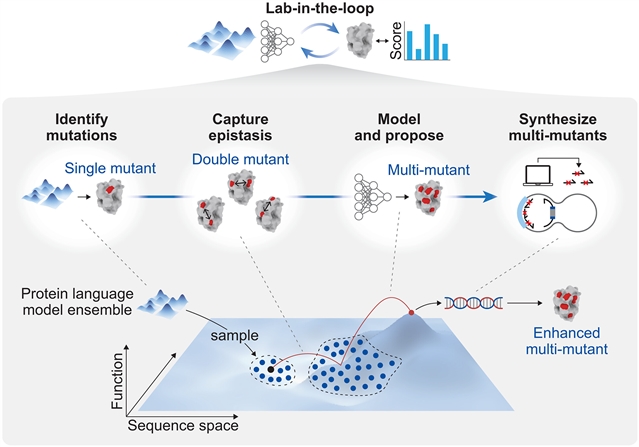
该课题组人员提出MULTI- evolution(MULTI代表模型引导的,通用的,有针对性的多突变体安装),这是一个系统地设计多突变体的快速进化框架。他们的方法将蛋白质语言模型或现有的功能数据与上位建模相结合,以预测协同组合。提出的多突变体是通过MULTI-assembly构建的,MULTI-assembly是一种能够在多碱基序列之间高效组装的诱变方法。将MULTI-evolution应用于三种蛋白质,通过一轮机器学习引导的定向进化,实现了高达10倍的改进。MULTI-evolution为广泛的蛋白质类型和功能提供了端到端多突变工程的简化方法。
研究人员表示,在高维序列空间中寻找协同突变组合的效率低下,限制了蛋白质工程的发展。传统方法的主题是逐步突变叠加,而机器学习方法需要大量的数据集或多轮实验,并且受到成本高、长度有限的基因合成的瓶颈。
附:英文原文
Title: Rapid directed evolution guided by protein language models and epistatic interactions
Author: Vincent Q. Tran, Matthew Nemeth, Liam J. Bartie, Sita S. Chandrasekaran, Alison Fanton, Hyungseok C. Moon, Brian L. Hie, Silvana Konermann, Patrick D. Hsu
Issue&Volume: 2026-05-07
Abstract: Protein engineering is limited by the inefficient search through a high-dimensional sequence space to find combinations of synergistic mutations. Traditional approaches use stepwise mutation stacking, whereas machine learning methods require extensive datasets or multiple experimental rounds and are bottlenecked by costly, length-limited gene synthesis. We present MULTI-evolve (where MULTI stands for model-guided, universal, targeted installation of multimutants), a rapid evolution framework that systematically engineers multimutants. Our approach combines protein language models or existing functional data with epistatic modeling to predict synergistic combinations. Proposed multimutants are built through MULTI-assembly, a mutagenesis method enabling high-efficiency assembly across multikilobase sequences. Applying MULTI-evolve to three proteins achieved up to 10-fold improvements with a single round of machine learning–guided directed evolution. MULTI-evolve provides a streamlined approach for end-to-end, multimutant engineering for a broad range of protein types and functions.
DOI: aea1820
Source: https://www.science.org/doi/10.1126/science.aea1820
