加州理工学院William M. Clemons Jr.团队宣布他们研制了噬菌体裂解蛋白抑制聚合MurJ翻转酶。这一研究成果发表在2026年2月25日出版的国际学术期刊《自然》上。
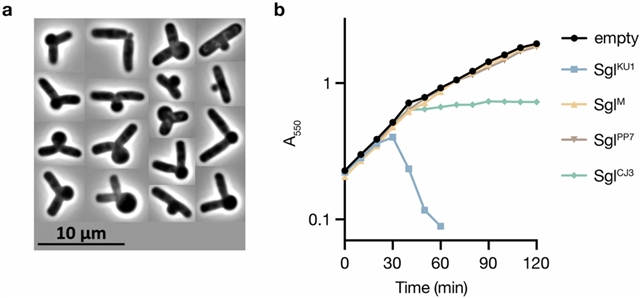
在这里,小组描述了这些噬菌体编码的Sgls抑制MurJ的共同机制。研究组确定了结合MurJ的SglM和SglPP7的结构,并发现了与预测的噬菌体Changjiang3 (SglCJ3)不同的第三个靶向MurJ的Sgl,研究组也对其进行了结构表征。他们的研究结果表明,所有三种Sgls都通过共同的MurJ界面趋同地将MurJ捕获在外质开放构象中,从而揭示了药物设计的途径。
据了解,抗微生物药物耐药性是一项全球性的健康挑战,需要确定新的可药物靶点。必需脂质II翻转酶MurJ是细菌细胞壁生物合成中一个很有前景但尚未被充分开发的抗菌靶点。唯一已知的革兰氏阴性(diderm) MurJ抑制剂是来自裂解单链RNA噬菌体M (SglM)和PP7 (SglPP7)的单基因裂解蛋白(Sgls)。SglM和SglPP7具有不同的进化起源,没有序列相似性。
附:英文原文
Title: Convergent MurJ flippase inhibition by phage lysis proteins
Author: Li, Yancheng E., Antillon, S. Francesca, Baron, Grace F., Chamakura, Karthik, Young, Ry, Clemons, William M.
Issue&Volume: 2026-02-25
Abstract: Antimicrobial drug resistance poses a global health challenge that necessitates the identification of new druggable targets1,2,3. The essential lipidII flippase MurJ is a promising yet underexplored antimicrobial target in bacterial cell wall biosynthesis4,5,6,7. The only known inhibitors of Gram-negative (diderm) MurJ are the single-gene lysis proteins (Sgls) from the lytic single-strand RNA phages M (SglM) and PP7 (SglPP7)8,9. SglM and SglPP7 have distinct evolutionary origins and share no sequence similarity. Here we describe a common mechanism of MurJ inhibition by these phage-encoded Sgls. We determined the structures of MurJ-bound SglM and SglPP7 and discovered a third distinct MurJ-targeting Sgl from the predicted phage Changjiang3 (SglCJ3) that we also characterized structurally. Our findings demonstrate that all three Sgls evolved convergently to trap MurJ in a periplasm-open conformation through a common MurJ interface, revealing a pathway for drug design.
DOI: 10.1038/s41586-026-10163-w
Source: https://www.nature.com/articles/s41586-026-10163-w
Nature:《自然》,创刊于1869年。隶属于施普林格·自然出版集团,最新IF:69.504
官方网址:http://www.nature.com/
投稿链接:http://www.nature.com/authors/submit_manuscript.html
